4932438A13Rik
Mus musculus RIKEN cDNA 4932438A13 gene (4932438A13Rik)
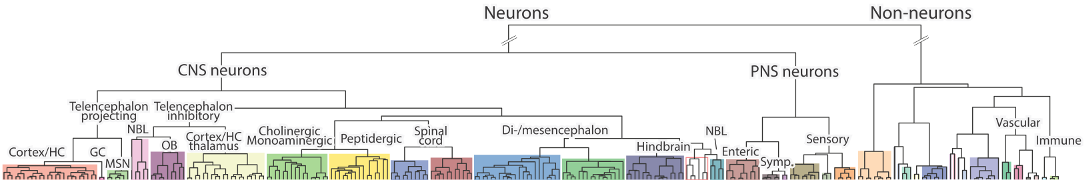

Clusters expressing 4932438A13Rik
Table shows clusters that express the gene at a trinarization score >= 0.95
| Expression | Index | Symbol | Description | Region |
| 0.3317 | 2 | TEGLU3 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.438 | 3 | TEGLU2 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.5091 | 4 | TEGLU20 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.4201 | 5 | TEGLU11 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.4223 | 6 | TEGLU12 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.6409 | 7 | TEGLU10 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.4208 | 8 | TEGLU9 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.3562 | 9 | TEGLU8 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.3663 | 10 | TEGLU7 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.4646 | 12 | TEGLU13 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.8857 | 13 | TEGLU14 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.4683 | 14 | TEGLU5 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.6393 | 15 | TEGLU16 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.6882 | 16 | TEGLU15 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.4923 | 17 | TEGLU17 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.4429 | 18 | TEGLU18 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.3708 | 19 | TEGLU19 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.38 | 20 | TEGLU22 | Excitatory neurons, amygdala | Telencephalon |
| 0.7596 | 21 | TEGLU21 | Excitatory neurons, hippocampus CA1 | Telencephalon |
| 0.5527 | 22 | TEGLU4 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.3098 | 23 | TEGLU24 | Excitatory neurons, hippocampus CA1 | Telencephalon |
| 0.4855 | 24 | TEGLU23 | Excitatory neurons, hippocampus CA3 | Telencephalon |
| 0.336 | 27 | MSN1 | D1 medium spiny neurons, striatum | Telencephalon |
| 0.2697 | 28 | MSN2 | D2 medium spiny neurons, striatum | Telencephalon |
| 0.2715 | 30 | MSN4 | D1 medium spiny neurons, striatum | Telencephalon |
| 0.3746 | 47 | DEINH1 | Inhibitory neurons, thalamus | Diencephalon |
| 0.3533 | 48 | DEINH2 | Inhibitory neurons, thalamus | Diencephalon |
| 0.3363 | 49 | TEINH17 | Axo-axonic, cortex/hippocampus | Telencephalon |
| 0.3419 | 50 | TEINH18 | Basket and bistratified cells, cortex/hippocampus | Telencephalon |
| 0.3712 | 51 | TEINH19 | Hippocamposeptal projection, cortex/hippocampus | Telencephalon |
| 0.3043 | 53 | TEINH16 | Ivy and MGE-derived neurogliaform cells, cortex/hippocampus | Telencephalon |
| 0.3957 | 56 | TEINH20 | Inhibitory interneurons, hippocampus | Telencephalon |
| 0.5767 | 57 | TEINH13 | Trilaminar cells, hippocampus | Telencephalon |
| 0.5857 | 58 | TEINH12 | Non-border Cck interneurons, cortex/hippocampus | Telencephalon |
| 0.7045 | 61 | TEINH11 | R-LM border Cck interneurons, cortex/hippocampus | Telencephalon |
| 0.7438 | 67 | TECHO | Cholinergic interneurons, telencephalon | Telencephalon |
| 0.8333 | 68 | DECHO1 | Cholinergic neurons, septal nucleus, Meissnert and diagonal band | Diencephalon |
| 1.0577 | 69 | HBCHO4 | Afferent nuclei of cranial nerves III-V | Rhombencephalon |
| 1.9167 | 70 | HBCHO3 | Afferent nuclei of cranial nerves VI-XII | Rhombencephalon |
| 0.675 | 71 | HBADR | Adrenergic cell groups of the medulla | Rhombencephalon |
| 0.4783 | 72 | HBNOR | Noradrenergic neurons of the medulla | Rhombencephalon |
| 0.5957 | 75 | MEGLU14 | Glutamatergic projection neurons of the raphe nucleus | Mesencephalon |
| 0.4286 | 77 | MBDOP2 | Dopaminergic neurons, ventral midbrain (SNc, VTA) | Mesencephalon |
| 0.6762 | 78 | HBSER1 | Serotonergic neurons, hindbrain | Rhombencephalon |
| 1.3119 | 79 | HBSER2 | Serotonergic neurons, hindbrain | Rhombencephalon |
| 0.6437 | 80 | HBSER3 | Serotonergic neurons, hindbrain | Rhombencephalon |
| 1.4531 | 81 | HBSER5 | Serotonergic neurons, hindbrain | Rhombencephalon |
| 1.0694 | 82 | HBSER4 | Serotonergic neurons, hindbrain | Rhombencephalon |
| 0.3172 | 83 | TEINH3 | Inhibitory neurons, telencephalon | Telencephalon |
| 0.5102 | 106 | HBINH9 | Inhibitory neurons, hindbrain | Rhombencephalon |
| 0.4486 | 119 | HBGLU3 | Excitatory neurons, hindbrain | Rhombencephalon |
| 0.695 | 120 | HBGLU2 | Excitatory neurons, hindbrain | Rhombencephalon |
| 0.3482 | 121 | MEGLU2 | Excitatory neurons, midbrain | Mesencephalon |
| 0.3028 | 123 | DEGLU5 | Excitatory neurons, midbrain | Mesencephalon |
| 0.4411 | 124 | MEGLU1 | Excitatory neurons, midbrain | Mesencephalon |
| 0.2885 | 126 | MEGLU8 | Excitatory neurons, midbrain | Mesencephalon |
| 0.2768 | 128 | MEGLU10 | Excitatory neurons, midbrain | Mesencephalon |
| 0.3735 | 129 | MEGLU11 | Excitatory neurons, midbrain | Mesencephalon |
| 0.4677 | 130 | MBCHO1 | Cholinergic neurons, midbrain red nucleus | Mesencephalon |
| 0.7063 | 131 | MEGLU6 | Excitatory neurons, midbrain | Mesencephalon |
| 0.3452 | 132 | MEGLU5 | Excitatory neurons, midbrain | Mesencephalon |
| 0.3015 | 133 | MEGLU4 | Excitatory neurons, midbrain | Mesencephalon |
| 0.9423 | 136 | HBGLU1 | Excitatory neurons, hindbrain | Rhombencephalon |
| 0.69 | 137 | DEGLU1 | Excitatory neurons, thalamus | Diencephalon |
| 0.7704 | 138 | DEGLU2 | Excitatory neurons, hypothalamus | Diencephalon |
| 0.5894 | 139 | DEGLU3 | Excitatory neurons, thalamus | Diencephalon |
| 0.495 | 140 | DEGLU4 | Excitatory neurons, thalamus | Diencephalon |
| 0.3673 | 141 | MEINH12 | Inhibitory neurons, midbrain | Mesencephalon |
| 0.327 | 143 | MEINH10 | Inhibitory neurons, midbrain | Mesencephalon |
| 0.2575 | 144 | MEINH9 | Inhibitory neurons, midbrain | Mesencephalon |
| 0.4935 | 145 | MEINH5 | Inhibitory neurons, midbrain | Mesencephalon |
| 0.4894 | 146 | MEINH6 | Inhibitory neurons, midbrain | Mesencephalon |
| 0.4198 | 148 | MEINH4 | Inhibitory neurons, midbrain | Mesencephalon |
| 0.3721 | 149 | MEINH3 | Inhibitory neurons, midbrain | Mesencephalon |
| 0.7833 | 150 | HBINH5 | Inhibitory neurons, hindbrain | Rhombencephalon |
| 0.4512 | 151 | MEINH2 | Inhibitory neurons, midbrain | Mesencephalon |
| 0.4211 | 152 | DEINH3 | Inhibitory neurons, hypothalamus | Diencephalon |
| 0.4219 | 153 | TEINH1 | Inhibitory neurons, pallidum | Telencephalon |
| 0.4615 | 156 | HBINH1 | Inhibitory neurons, hindbrain | Rhombencephalon |
| 0.4133 | 158 | HBINH4 | Inhibitory neurons, hindbrain | Rhombencephalon |
| 0.7903 | 159 | HBINH6 | Inhibitory neurons, hindbrain | Rhombencephalon |
| 1.5909 | 160 | HBINH2 | Inhibitory neurons, hindbrain | Rhombencephalon |
| 2.0 | 161 | HBCHO1 | Cholinergic neurons, hindbrain | Rhombencephalon |
| 0.7727 | 162 | HBCHO2 | Cholinergic neurons, hindbrain | Rhombencephalon |
| 0.7143 | 163 | HBGLU4 | Excitatory neurons, hindbrain | Rhombencephalon |
| 0.5154 | 164 | HBGLU5 | Excitatory neurons, hindbrain | Rhombencephalon |
| 1.1071 | 165 | HBGLU6 | Excitatory neurons, hindbrain | Rhombencephalon |
| 1.04 | 166 | HBGLU7 | Excitatory neurons, hindbrain | Rhombencephalon |
| 2.0909 | 167 | HBGLU8 | Excitatory neurons, hindbrain | Rhombencephalon |
| 0.4536 | 168 | HBGLU9 | Excitatory neurons, hindbrain | Rhombencephalon |
| 0.52 | 169 | HBINH7 | Inhibitory neurons, hindbrain | Rhombencephalon |
| 0.4545 | 170 | HBINH8 | Inhibitory neurons, hindbrain | Rhombencephalon |
| 0.3237 | 180 | OBNBL1 | Neuroblasts, olfactory | Telencephalon |
| 0.8571 | 182 | ENT1 | Nitrergic enteric neurons | Neural crest |
| 0.7383 | 183 | ENT2 | Nitrergic enteric neurons | Neural crest |
| 0.529 | 184 | ENT3 | Nitrergic enteric neurons | Neural crest |
| 0.3935 | 185 | ENT4 | Cholinergic enteric neurons | Neural crest |
| 0.4746 | 187 | ENT6 | Cholinergic enteric neurons | Neural crest |
| 0.4839 | 188 | ENT7 | Cholinergic enteric neurons, VGLUT2 | Neural crest |
| 0.7835 | 189 | ENT8 | Cholinergic enteric neurons, VGLUT2 | Neural crest |
| 1.0118 | 190 | ENT9 | Cholinergic enteric neurons | Neural crest |
| 0.3301 | 193 | SYNOR3 | Noradrenergic neurons, sympathetic | Neural crest |
| 0.4176 | 194 | SYNOR4 | Noradrenergic erector muscle neurons | Neural crest |
| 0.413 | 195 | SYNOR5 | Noradrenergic erector muscle neurons | Neural crest |
| 0.44 | 196 | SYCHO2 | Cholinergic neurons, sympathetic | Neural crest |
| 0.9333 | 197 | SYCHO1 | Cholinergic neurons, sympathetic | Neural crest |
| 0.4216 | 202 | PSPEP2 | Peptidergic (PEP1.3), DRG | Neural crest |
| 0.3504 | 203 | PSPEP4 | Peptidergic (PEP1.1), DRG | Neural crest |
| 0.5978 | 204 | PSPEP3 | Peptidergic (PEP1..4), DRG | Neural crest |
| 0.4474 | 208 | PSNF1 | Neurofilament (NF1), DRG | Neural crest |
| 0.4113 | 209 | PSNP1 | Non-peptidergic (TH), DRG | Neural crest |
| 0.7083 | 210 | PSNP2 | Non-peptidergic (NP1.1), DRG | Neural crest |
| 0.6556 | 211 | PSNP3 | Non-peptidergic (NP1.2), DRG | Neural crest |
| 0.9333 | 212 | PSNP4 | Non-peptidergic (NP2.1), DRG | Neural crest |
| 0.375 | 213 | PSNP5 | Non-peptidergic (NP2.2), DRG | Neural crest |
| 0.553 | 214 | PSNP6 | Non-peptidergic (NP3), DRG | Neural crest |
| 0.2535 | 220 | MFOL1 | Myelin forming oligodendrocytes (MFOL) | CNS |
4932438A13Rik is a marker of the following clusters
| Expression | Index | Symbol | Description | Region |