9530068E07Rik
Mus musculus RIKEN cDNA 9530068E07 gene (9530068E07Rik)
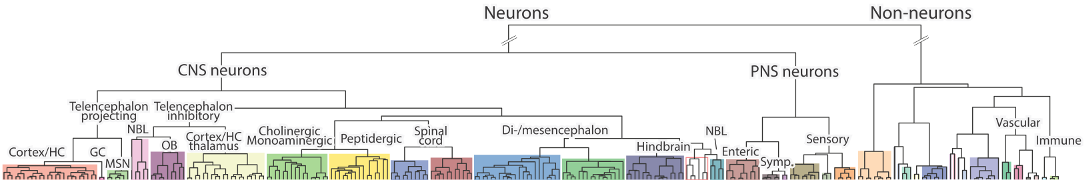

Clusters expressing 9530068E07Rik
Table shows clusters that express the gene at a trinarization score >= 0.95
| Expression | Index | Symbol | Description | Region |
| 0.3978 | 3 | TEGLU2 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.4909 | 4 | TEGLU20 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.3641 | 5 | TEGLU11 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.3368 | 6 | TEGLU12 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.4619 | 7 | TEGLU10 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.3564 | 8 | TEGLU9 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.3406 | 9 | TEGLU8 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.4238 | 10 | TEGLU7 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.4961 | 12 | TEGLU13 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.6857 | 13 | TEGLU14 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.3889 | 14 | TEGLU5 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.5161 | 16 | TEGLU15 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.4273 | 17 | TEGLU17 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.4658 | 18 | TEGLU18 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.4045 | 19 | TEGLU19 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.3424 | 20 | TEGLU22 | Excitatory neurons, amygdala | Telencephalon |
| 0.5577 | 21 | TEGLU21 | Excitatory neurons, hippocampus CA1 | Telencephalon |
| 0.5286 | 22 | TEGLU4 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.3814 | 23 | TEGLU24 | Excitatory neurons, hippocampus CA1 | Telencephalon |
| 0.5496 | 24 | TEGLU23 | Excitatory neurons, hippocampus CA3 | Telencephalon |
| 0.3072 | 27 | MSN1 | D1 medium spiny neurons, striatum | Telencephalon |
| 0.2514 | 28 | MSN2 | D2 medium spiny neurons, striatum | Telencephalon |
| 0.3304 | 47 | DEINH1 | Inhibitory neurons, thalamus | Diencephalon |
| 0.3185 | 49 | TEINH17 | Axo-axonic, cortex/hippocampus | Telencephalon |
| 0.4369 | 50 | TEINH18 | Basket and bistratified cells, cortex/hippocampus | Telencephalon |
| 0.4548 | 51 | TEINH19 | Hippocamposeptal projection, cortex/hippocampus | Telencephalon |
| 0.3636 | 52 | TEINH21 | Sleep-active, long-range projection interneurons, cortex/hippocampus | Telencephalon |
| 0.3433 | 54 | TEINH15 | CGE-derived neurogliaform cells, cortex/hippocampus | Telencephalon |
| 0.5612 | 56 | TEINH20 | Inhibitory interneurons, hippocampus | Telencephalon |
| 0.589 | 57 | TEINH13 | Trilaminar cells, hippocampus | Telencephalon |
| 0.6143 | 58 | TEINH12 | Non-border Cck interneurons, cortex/hippocampus | Telencephalon |
| 0.4074 | 59 | TEINH9 | Non-border Cck interneurons, hippocampus | Telencephalon |
| 0.4389 | 60 | TEINH10 | R-LM border Cck interneurons, cortex/hippocampus | Telencephalon |
| 0.5833 | 61 | TEINH11 | R-LM border Cck interneurons, cortex/hippocampus | Telencephalon |
| 0.4118 | 65 | TEINH7 | Interneuron-selective interneurons, hippocampus | Telencephalon |
| 0.7438 | 67 | TECHO | Cholinergic interneurons, telencephalon | Telencephalon |
| 0.7456 | 68 | DECHO1 | Cholinergic neurons, septal nucleus, Meissnert and diagonal band | Diencephalon |
| 1.3077 | 69 | HBCHO4 | Afferent nuclei of cranial nerves III-V | Rhombencephalon |
| 1.7778 | 70 | HBCHO3 | Afferent nuclei of cranial nerves VI-XII | Rhombencephalon |
| 0.55 | 71 | HBADR | Adrenergic cell groups of the medulla | Rhombencephalon |
| 0.4783 | 72 | HBNOR | Noradrenergic neurons of the medulla | Rhombencephalon |
| 1.0938 | 74 | HYPEP6 | Orexin-producing neurons, hypothalamus | Diencephalon |
| 1.0213 | 75 | MEGLU14 | Glutamatergic projection neurons of the raphe nucleus | Mesencephalon |
| 0.3509 | 76 | MBDOP1 | Dopaminergic neurons, periaqueductal grey | Mesencephalon |
| 0.873 | 77 | MBDOP2 | Dopaminergic neurons, ventral midbrain (SNc, VTA) | Mesencephalon |
| 1.1238 | 78 | HBSER1 | Serotonergic neurons, hindbrain | Rhombencephalon |
| 1.0826 | 79 | HBSER2 | Serotonergic neurons, hindbrain | Rhombencephalon |
| 0.8391 | 80 | HBSER3 | Serotonergic neurons, hindbrain | Rhombencephalon |
| 0.9531 | 81 | HBSER5 | Serotonergic neurons, hindbrain | Rhombencephalon |
| 0.9167 | 82 | HBSER4 | Serotonergic neurons, hindbrain | Rhombencephalon |
| 0.6207 | 83 | TEINH3 | Inhibitory neurons, telencephalon | Telencephalon |
| 0.3701 | 86 | DEINH5 | Peptidergic neurons, hypothalamus | Diencephalon |
| 0.4079 | 87 | HYPEP3 | Peptidergic neurons, hypothalamus | Diencephalon |
| 0.4241 | 88 | HYPEP1 | Peptidergic neurons, hypothalamus | Diencephalon |
| 0.4651 | 89 | HYPEP2 | Peptidergic neurons, hypothalamus | Diencephalon |
| 0.9167 | 91 | DEINH6 | Peptidergic neurons, hypothalamus | Diencephalon |
| 0.474 | 94 | HYPEP5 | Vasopressin-producing cells, hypothalamus | Diencephalon |
| 0.3846 | 98 | SCINH10 | Inhibitory neurons, spinal cord | Spinal cord |
| 0.3333 | 102 | SCINH6 | Inhibitory neurons, spinal cord | Spinal cord |
| 0.515 | 120 | HBGLU2 | Excitatory neurons, hindbrain | Rhombencephalon |
| 0.3601 | 121 | MEGLU2 | Excitatory neurons, midbrain | Mesencephalon |
| 0.3431 | 122 | MEGLU3 | Excitatory neurons, midbrain | Mesencephalon |
| 0.3451 | 123 | DEGLU5 | Excitatory neurons, midbrain | Mesencephalon |
| 0.4335 | 124 | MEGLU1 | Excitatory neurons, midbrain | Mesencephalon |
| 0.2826 | 125 | MEGLU7 | Excitatory neurons, midbrain | Mesencephalon |
| 0.4156 | 127 | MEGLU9 | Excitatory neurons, midbrain | Mesencephalon |
| 0.3437 | 128 | MEGLU10 | Excitatory neurons, midbrain | Mesencephalon |
| 0.3574 | 129 | MEGLU11 | Excitatory neurons, midbrain | Mesencephalon |
| 0.5806 | 130 | MBCHO1 | Cholinergic neurons, midbrain red nucleus | Mesencephalon |
| 0.5175 | 131 | MEGLU6 | Excitatory neurons, midbrain | Mesencephalon |
| 0.3025 | 132 | MEGLU5 | Excitatory neurons, midbrain | Mesencephalon |
| 0.6154 | 136 | HBGLU1 | Excitatory neurons, hindbrain | Rhombencephalon |
| 0.6361 | 137 | DEGLU1 | Excitatory neurons, thalamus | Diencephalon |
| 0.5136 | 138 | DEGLU2 | Excitatory neurons, hypothalamus | Diencephalon |
| 0.4178 | 139 | DEGLU3 | Excitatory neurons, thalamus | Diencephalon |
| 0.5644 | 140 | DEGLU4 | Excitatory neurons, thalamus | Diencephalon |
| 0.2992 | 143 | MEINH10 | Inhibitory neurons, midbrain | Mesencephalon |
| 0.3571 | 145 | MEINH5 | Inhibitory neurons, midbrain | Mesencephalon |
| 0.4255 | 146 | MEINH6 | Inhibitory neurons, midbrain | Mesencephalon |
| 0.5494 | 148 | MEINH4 | Inhibitory neurons, midbrain | Mesencephalon |
| 0.5233 | 149 | MEINH3 | Inhibitory neurons, midbrain | Mesencephalon |
| 0.5583 | 150 | HBINH5 | Inhibitory neurons, hindbrain | Rhombencephalon |
| 0.6356 | 151 | MEINH2 | Inhibitory neurons, midbrain | Mesencephalon |
| 0.548 | 152 | DEINH3 | Inhibitory neurons, hypothalamus | Diencephalon |
| 0.5417 | 153 | TEINH1 | Inhibitory neurons, pallidum | Telencephalon |
| 0.5857 | 154 | MEINH13 | Inhibitory neurons, midbrain | Mesencephalon |
| 0.6129 | 159 | HBINH6 | Inhibitory neurons, hindbrain | Rhombencephalon |
| 0.9545 | 160 | HBINH2 | Inhibitory neurons, hindbrain | Rhombencephalon |
| 0.9 | 161 | HBCHO1 | Cholinergic neurons, hindbrain | Rhombencephalon |
| 0.9091 | 162 | HBCHO2 | Cholinergic neurons, hindbrain | Rhombencephalon |
| 1.0 | 163 | HBGLU4 | Excitatory neurons, hindbrain | Rhombencephalon |
| 0.8923 | 164 | HBGLU5 | Excitatory neurons, hindbrain | Rhombencephalon |
| 0.6548 | 165 | HBGLU6 | Excitatory neurons, hindbrain | Rhombencephalon |
| 0.7345 | 166 | HBGLU7 | Excitatory neurons, hindbrain | Rhombencephalon |
| 0.9091 | 167 | HBGLU8 | Excitatory neurons, hindbrain | Rhombencephalon |
| 0.6392 | 168 | HBGLU9 | Excitatory neurons, hindbrain | Rhombencephalon |
| 0.4 | 169 | HBINH7 | Inhibitory neurons, hindbrain | Rhombencephalon |
| 0.4245 | 180 | OBNBL1 | Neuroblasts, olfactory | Telencephalon |
| 1.2857 | 182 | ENT1 | Nitrergic enteric neurons | Neural crest |
| 1.1589 | 183 | ENT2 | Nitrergic enteric neurons | Neural crest |
| 0.8645 | 184 | ENT3 | Nitrergic enteric neurons | Neural crest |
| 1.0452 | 185 | ENT4 | Cholinergic enteric neurons | Neural crest |
| 0.4595 | 186 | ENT5 | Cholinergic enteric neurons | Neural crest |
| 1.0254 | 187 | ENT6 | Cholinergic enteric neurons | Neural crest |
| 1.4194 | 188 | ENT7 | Cholinergic enteric neurons, VGLUT2 | Neural crest |
| 1.4742 | 189 | ENT8 | Cholinergic enteric neurons, VGLUT2 | Neural crest |
| 1.4083 | 190 | ENT9 | Cholinergic enteric neurons | Neural crest |
| 1.5052 | 191 | SYNOR1 | Noradrenergic erector muscle neurons | Neural crest |
| 1.3065 | 192 | SYNOR2 | Noradrenergic neurons, sympathetic | Neural crest |
| 1.4401 | 193 | SYNOR3 | Noradrenergic neurons, sympathetic | Neural crest |
| 1.6154 | 194 | SYNOR4 | Noradrenergic erector muscle neurons | Neural crest |
| 1.1087 | 195 | SYNOR5 | Noradrenergic erector muscle neurons | Neural crest |
| 1.4533 | 196 | SYCHO2 | Cholinergic neurons, sympathetic | Neural crest |
| 1.7333 | 197 | SYCHO1 | Cholinergic neurons, sympathetic | Neural crest |
| 0.7857 | 198 | PSPEP8 | Peptidergic (TrpM8), DRG | Neural crest |
| 0.2813 | 199 | PSPEP7 | Peptidergic (TrpM8), DRG | Neural crest |
| 0.5 | 200 | PSPEP6 | Peptidergic (TrpM8), DRG | Neural crest |
| 0.4333 | 201 | PSPEP5 | Peptidergic (PEP1.2), DRG | Neural crest |
| 0.951 | 202 | PSPEP2 | Peptidergic (PEP1.3), DRG | Neural crest |
| 0.6752 | 203 | PSPEP4 | Peptidergic (PEP1.1), DRG | Neural crest |
| 0.837 | 204 | PSPEP3 | Peptidergic (PEP1..4), DRG | Neural crest |
| 0.9744 | 205 | PSPEP1 | Peptidergicv (PEP2), DRG | Neural crest |
| 1.7419 | 206 | PSNF3 | Neurofilament (NF2/3), DRG | Neural crest |
| 1.5088 | 207 | PSNF2 | Neurofilament (NF4/5), DRG | Neural crest |
| 0.8684 | 208 | PSNF1 | Neurofilament (NF1), DRG | Neural crest |
| 1.0071 | 209 | PSNP1 | Non-peptidergic (TH), DRG | Neural crest |
| 1.1944 | 210 | PSNP2 | Non-peptidergic (NP1.1), DRG | Neural crest |
| 0.8815 | 211 | PSNP3 | Non-peptidergic (NP1.2), DRG | Neural crest |
| 0.9333 | 212 | PSNP4 | Non-peptidergic (NP2.1), DRG | Neural crest |
| 0.8977 | 213 | PSNP5 | Non-peptidergic (NP2.2), DRG | Neural crest |
| 1.0 | 214 | PSNP6 | Non-peptidergic (NP3), DRG | Neural crest |
| 0.2779 | 218 | NFOL1 | Newly formed oligodendrocytes (NFOL) | CNS |
| 0.3031 | 219 | MFOL2 | Myelin forming oligodendrocytes (MFOL) | CNS |
| 0.2512 | 222 | MOL2 | Mature oligodendrocytes, hindbrain | CNS |
| 0.3959 | 223 | MOL3 | Mature oligodendrocytes, spinal cord enriched (high Klk6) | CNS |
| 0.714 | 224 | CHOR | Chorid plexus epithelial cells | CNS |
| 0.4595 | 225 | HYPEN | Hypendymal cell, subcommissural organ | CNS |
| 0.5169 | 226 | EPSC | Ependymal cells, spinal cord | CNS |
| 0.9587 | 227 | EPEN | Ependymal cells | CNS |
| 0.4681 | 240 | SCHW | Schwann cells | Neural crest |
| 0.4725 | 241 | SATG2 | Satellite glia | Neural crest |
| 0.4667 | 242 | SATG1 | Satellite glia, proliferating | Neural crest |
| 1.5276 | 243 | ENTG1 | Enteric glia, proliferating | Neural crest |
| 0.5657 | 244 | ENTG2 | Enteric glia | Neural crest |
| 0.9595 | 245 | ENTG3 | Enteric glia | Neural crest |
| 0.8185 | 246 | ENTG4 | Enteric glia | Neural crest |
| 0.8004 | 247 | ENTG5 | Enteric glia | Neural crest |
| 0.9038 | 248 | ENTG6 | Enteric glia | Neural crest |
| 0.9183 | 249 | ENTG7 | Enteric glia | Neural crest |
| 1.2449 | 250 | ENMFB | Enteric mesothelial fibroblasts | Neural crest |
| 0.7956 | 251 | ABC | Vascular leptomeningeal cells | Neural crest |
| 0.3172 | 252 | VLMC2 | Vascular leptomeningeal cells | Neural crest |
| 0.3761 | 254 | VECA | Vascular endothelial cells, arterial | Mesoderm |
| 0.3202 | 257 | PER1 | Pericytes | Mesoderm |
| 0.5082 | 258 | PER2 | Pericytes, possibly mixed with VENC | Mesoderm |
| 0.3015 | 259 | VECC | Vascular endothelial cells, capillary | Mesoderm |
9530068E07Rik is a marker of the following clusters
| Expression | Index | Symbol | Description | Region |